MICROBIOME SEQUENCING (16S + ITS)
A New Standard In Pathogen Detection
DNA sequencing is the next generation of greenhouse pathogen detection, offering improved insights into both the healthy and pathogenic microbes present in a sample. Through 16S/ITS sequencing we are able to identify over 5,000 bacterial and fungal species present in each sample, and determine what portion of the total community they make up. By determining the relative abundance of disease causing organisms in a sample, we can gain improved insights into which organisms are causing disease symptoms. This can help reduce confusion currently associated with interpreting PCR based testing results. Additionally, sequencing based testing allows us to test for all known pathogens in every sample instead of a small number of pre-selected species.

Pricing
Sample pricing includes:
Identification of +5,000 bacterial and fungal species per sample
Summary report detailing the most prevalent pathogens, bacteria, and fungi in each sample including information on how much of the community each species makes up
A brief description of each bacteria and fungal species mentioned in the summary report
A spreadsheet listing all of the species identified in each sample and their relative abundances
Support from our technical team to answer any questions you may have about your results
Download our comprehensive Disease Testing Sampling and Shipping Guide HERE
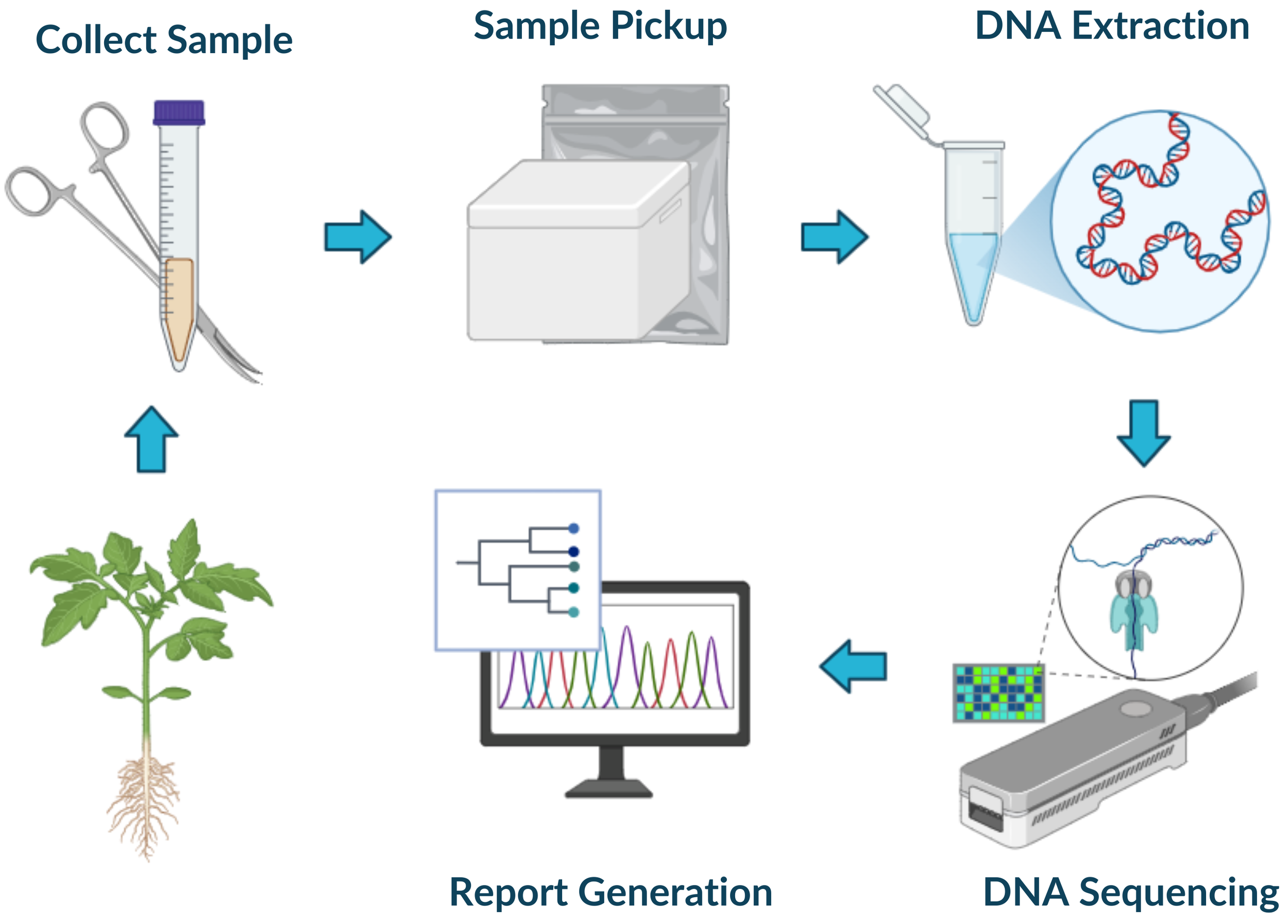
Our Workflow
Growers collect samples and place them inside a collection tube
Samples are either picked up by our team (Leamington / Kingsville) or shipped by the grower to our lab
DNA is extracted from the sample, the 16S and ITS genes are amplified, and the amplified DNA is purified
The sample is sequenced using long read DNA sequencing
The sequencing data is processed, the species within each sample are identified, and a report is generated
